Analysis: citizen scientists worldwide have come together to help design new drugs to take on the coronavirus
By staying local and social distancing, everyone is playing their part in stopping the spread of Covid-19, but what about playing a part in finding a treatment? If you have a computer, then you have everything you need to become a citizen scientist and take part in discovering new medicines for SARS-CoV-2.
Who are citizen scientists?
Citizen scientists are members of the public that volunteer to participate in a scientific project or activity. Current projects worldwide include the restoration of coral reefs in the Florida Keys and the discovery of new planets.
A good chunk of science requires a qualified person with a particular skill set. However, in the data collection stage, it usually comes down to sheer work-hours that may take a small group of researchers a long amount of time. Many hands make light work and retrieving scientific data is no exception.
Over the past few months, citizen scientists across the world have come together to design new drugs to treat coronavirus by playing a puzzle game. Others have produced the world's largest supercomputer to combat Covid-19 all from within their homes.
The colourful world of proteins
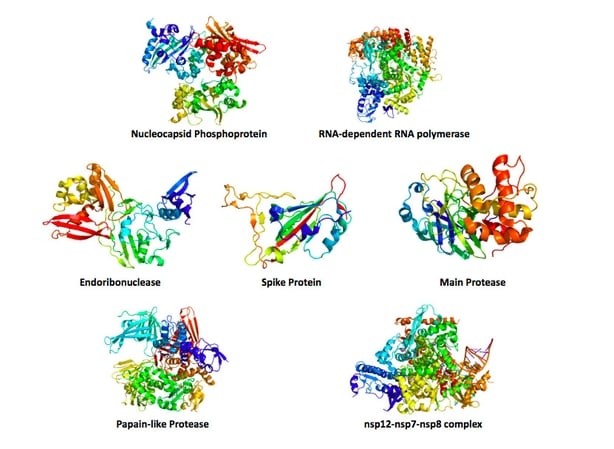
At first glance the picture above may look like something resembling a burst pinata, but they are in fact proteins. People normally think of proteins in a dietary context, but they are so much more than that post workout shake.
Proteins are the nanoscopic machines responsible for operating all life and there are thousands of different types working away in your body. Every protein forms a specific shape which gives it a specific function that can range anywhere from digesting food to moving our muscles.
While appearing harmless, the confetti bursts shown above are actually the key proteins that operate the coronavirus (not that colourful in real-life!). Just like a series of machines on a factory line, these proteins work together to allow the virus to attach, infect and replicate itself inside our cells.
We need your consent to load this rte-player contentWe use rte-player to manage extra content that can set cookies on your device and collect data about your activity. Please review their details and accept them to load the content.Manage Preferences
From RTÉ Radio 1's Drivetime, how the Environmental Protection Agency used Irish citizen scientists to monitor air quality
Our job as scientists is to design a spanner to throw into one of these machines to stop the virus from working (AKA a medicine). To make new medicines, we have to know and understand the protein we are targeting. Scientists can take a 3D snapshot of these proteins with relative ease using X-rays, but true understanding comes from seeing them in motion. By seeing how these tiny particles change their shape or ‘fold’, we can find weak spots that medicines can target to disable the virus.
The supercomputer in your gaff
We currently do not have the technology to observe proteins moving in real-time. Despite this hurdle, scientists can create computer simulations to predict how they move. The problem with making simulations is they require gigantic amounts of computing power and are generally done by state-of-the-art supercomputers.
But for those without a supercomputer lying about, this is problematic. Thinking outside the box, researchers from Stanford University developed a piece of software called Folding@home that uses distributed computing to create protein simulations. Instead of using one supercomputer to do one big job, distributed computing uses thousands of regular computers to do one small job each.
We need your consent to load this rte-player contentWe use rte-player to manage extra content that can set cookies on your device and collect data about your activity. Please review their details and accept them to load the content.Manage Preferences
From RTÉ Radio 1's Morning Ireland, meet Ireland's new €5.4 million supercomputer at the Irish Centre for High-End Computing
By downloading Folding@home, a user’s computer is sent a small calculation which it solves and sends back to the Folding@home server to then receive another. The joint effort of distributed computing can rival the power of supercomputers, provided a big enough community is created.
In March, Folding@home announced they were putting their efforts toward Covid-19 and the number of users exploded from 30,000 to 780,000 within a month, creating a network that was faster than the top 500 supercomputers in the world combined.
With our collective power, we are now at ~2.4 exaFLOPS (faster than the top 500 supercomputers combined)! We complement supercomputers like IBM Summit, which runs short calculations using 1000s of GPUs at once, by spreading longer calculations around the world in smaller chunks! pic.twitter.com/fdUaXOcdFJ
— Folding@home (@foldingathome) April 13, 2020
While researchers could target numerous different proteins to disable the coronavirus, one that stuck out was the Spike protein (no pun intended). This protein is the protrusion that gives the virus its infamous spikey shape and allows the virus to stick to human cells and infect us. Folding@home generated a simulation of this Spike protein that exposed a structural weak spot that could be used to develop an antiviral treatment for SARS-CoV-2.
After a few seconds into the simulation, the spike protein begins to reveal what resembles the opening of a monstrous mouth. Despite its non-monstrous size, this isn't far from the fact; it’s this mouth that the coronavirus uses to latch onto our cells and infect us. Covid-19 researchers have honed in on capping the spike protein as a potential approach to treat the coronavirus and Folding@home’s simulation has provided key information in how to do this.
While people may be apprehensive about folding@home slowing down their computer while they work, you can control the level of power folding@home uses and you can set it to only run when you aren’t using your computer. Interested in helping out? You can download Folding@home here
How video games are fighting Covid-19
Although computers are excellent at carrying out simple, step-wise commands, they lack imagination. This isn't particularly vital for making a financial report in Excel, but it plays an important role when you need to design a drug to fit into a 3D protein and people are excellent 3D puzzle solvers.
Inspired by this fact, collaborators at the University of Washington created a game called FoldIt that turns drug design into a simple, yet accurate puzzle game. One of the puzzles in this game involves the infamous Covid spike protein, the objective is to design a protein that complements and fits onto the spike protein. Creating such a protein could act as a plug that would jam the spike protein and render it useless. Disabling the spike protein would block it from attaching to human cells and stop the coronavirus from thriving within its host.
Despite the complex nature of drug development, Foldit elegantly simplifies protein design into an intuitive puzzle game while preserving the fundamentals surrounding this topic, enabling anyone to play the game. The greater your designed protein fits or 'binds' to the coronavirus spike protein, the higher the score.
Players who create the highest scoring proteins will have their designs made in the lab and tested against Covid-19. So far, 99 protein candidates have been selected and await testing. Previous to the Coronavirus outbreak, FoldIt players had solved protein structures that got them published in scientific journals. Who ever said playing video games was a waste of time?
The views expressed here are those of the author and do not represent or reflect the views of RTÉ